SMAP sliding-frames

This is the manual for the SMAP sliding-frames module of the SMAP package.
The first step prior to running SMAP haplotype-sites is the definition of the locus start and end points.
The module SMAP sliding-frames should be used to define sliding frames covering SNPs and/or structural variants in Shotgun data.
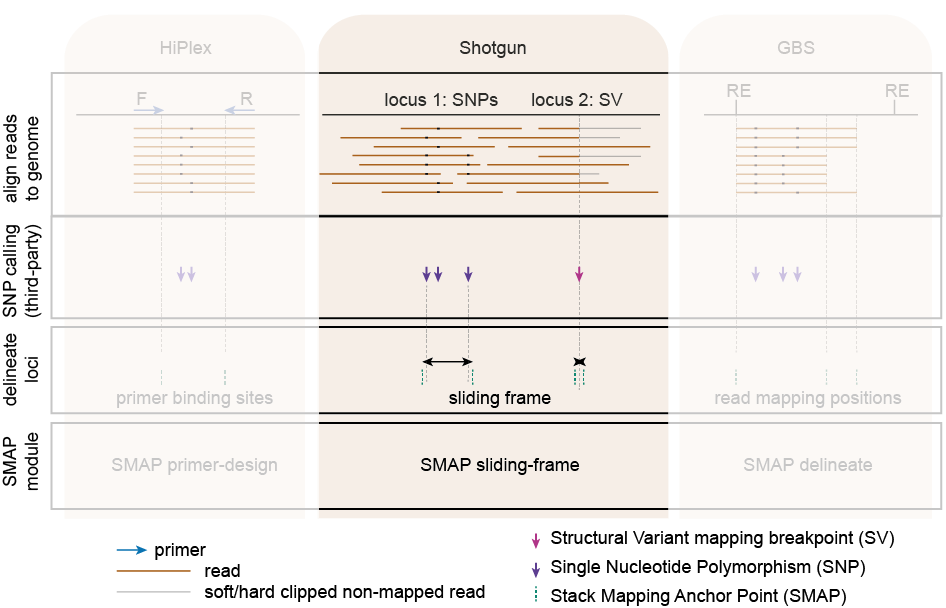
The module SMAP delineate should be run for GBS data to define relevant loci and read mapping polymorphisms in a data-driven manner.
The module SMAP design should be used for integrated design of HiPlex PCR primers and downstream analysis with SMAP haplotype-sites and/or SMAP haplotype-window.